Covariance functions
Exponential.RdFunctional form of covariance function assuming the argument is a
distance between locations. As they are defined here, they are in
fact correlation functions. To set the marginal variance (sill)
parameter, use the sigma argument in mKrig or Krig.
To set the nugget variance, use te tau2 argument in
mKrig or Krig.
Exponential(d, range = 1, alpha = 1/range, phi=1.0,theta = NULL)
Matern(d , range = 1,alpha=1/range, smoothness = 0.5,
nu= smoothness, phi=1.0)
Matern.cor.to.range(d, nu, cor.target=.5, guess=NULL,...)
RadialBasis(d,M,dimension, derivative = 0)Arguments
- d
Vector of distances or for
Matern.cor.to.rangejust a single distance.- range
Range parameter default is one. Note that the scale can also be specified through the "aRange" scaling argument used in fields covariance functions)
- alpha
1/range
- theta
Same as alpha
- phi
This parameter option is added to be compatible with older versions of fields and refers to the marginal variance of the process. e.g.
phi* exp( -d/aRange)is the exponential covariance for points separated by distance and range aRange. Throughout fields this parameter is equivalent to sigma and it recommended that sigma be used. If one is simulating random fields. See the help onsim.rffor more details.- smoothness
Smoothness parameter in Matern. Controls the number of derivatives in the process. Default is 1/2 corresponding to an exponential covariance.
- nu
Same as smoothness
- M
Interpreted as a spline M is the order of the derivatives in the penalty.
- dimension
Dimension of function
- cor.target
Correlation used to match the range parameter. Default is .5.
- guess
An optional starting guess for solution. This should not be needed.
- derivative
If greater than zero finds the first derivative of this function.
- ...
Additional arguments to pass to the bisection search function.
Details
Exponential:
exp( -d/range)
Matern:
con*(d**nu) * besselK(d , nu )
Matern covariance function transcribed from Stein's book page 31 nu==smoothness, alpha == 1/range
GeoR parameters map to kappa==smoothness and phi == range check for negative distances
con is a constant that normalizes the expression to be 1.0 when d=0.
Matern.cor.to.range:
This function is useful to find Matern covariance parameters that are
comparable for different smoothness parameters. Given a distance d,
smoothness nu, target correlation cor.target and
range aRange, this function determines numerically the value of
aRange so that
Matern( d, range=aRange, nu=nu) == cor.target
See the example for how this might be used.
Radial basis functions:
C.m,d r**(2m-d) d- odd
C.m,d r**(2m-d)ln(r) d-even
where C.m.d is a constant based on spline theory and r is the radial distance
between points. See radbas.constant for the computation of the constant.
Value
For the covariance functions: a vector of covariances.
For Matern.cor.to.range: the value of the range parameter.
References
Stein, M.L. (1999) Statistical Interpolation of Spatial Data: Some Theory for Kriging. Springer, New York.
See also
stationary.cov, stationary.image.cov, Wendland,stationary.taper.cov rad.cov
Examples
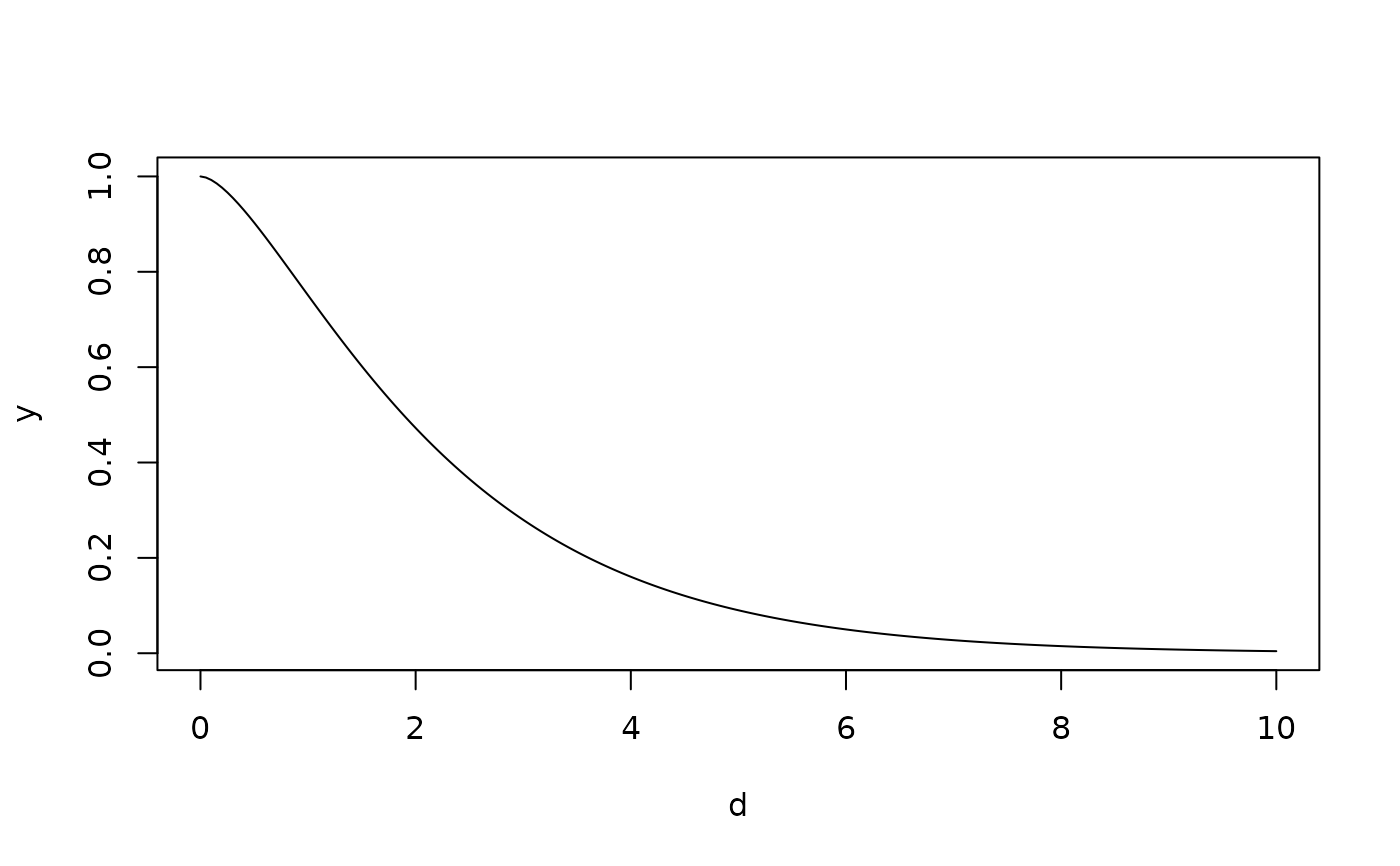
# a Matern correlation function
d<- seq( 0,10,,200)
y<- Matern( d, range=1.5, smoothness=1.0)
plot( d,y, type="l")
 # Several Materns of different smoothness with a similar correlation
# range
# find ranges for nu = .5, 1.0 and 2.0
# where the correlation drops to .1 at a distance of 10 units.
r1<- Matern.cor.to.range( 10, nu=.5, cor.target=.1)
r2<- Matern.cor.to.range( 10, nu=1.0, cor.target=.1)
r3<- Matern.cor.to.range( 10, nu=2.0, cor.target=.1)
# note that these equivalent ranges
# with respect to this correlation length are quite different
# due the different smoothness parameters.
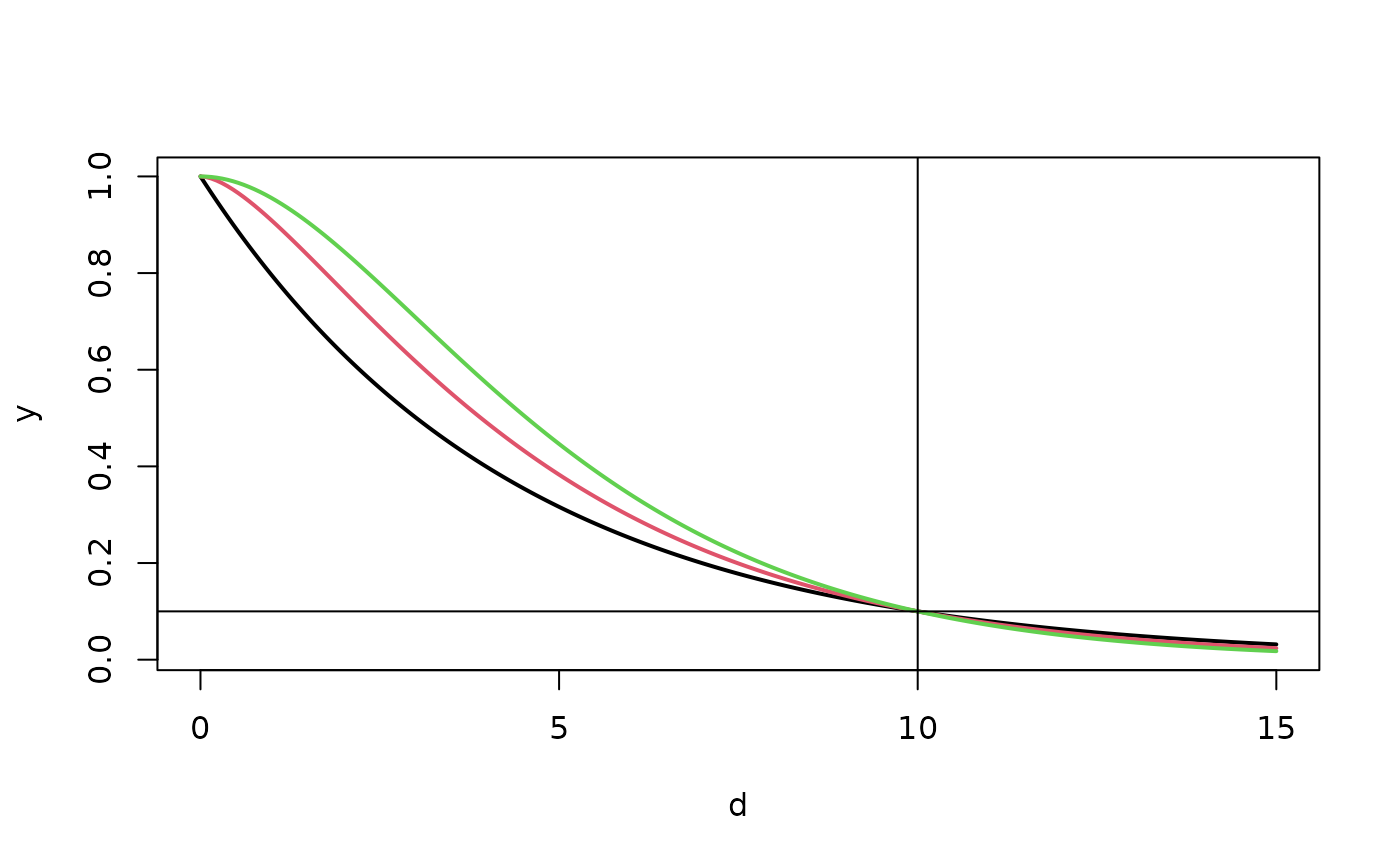
d<- seq( 0, 15,,200)
y<- cbind( Matern( d, range=r1, nu=.5),
Matern( d, range=r2, nu=1.0),
Matern( d, range=r3, nu=2.0))
matplot( d, y, type="l", lty=1, lwd=2)
xline( 10)
yline( .1)
# Several Materns of different smoothness with a similar correlation
# range
# find ranges for nu = .5, 1.0 and 2.0
# where the correlation drops to .1 at a distance of 10 units.
r1<- Matern.cor.to.range( 10, nu=.5, cor.target=.1)
r2<- Matern.cor.to.range( 10, nu=1.0, cor.target=.1)
r3<- Matern.cor.to.range( 10, nu=2.0, cor.target=.1)
# note that these equivalent ranges
# with respect to this correlation length are quite different
# due the different smoothness parameters.
d<- seq( 0, 15,,200)
y<- cbind( Matern( d, range=r1, nu=.5),
Matern( d, range=r2, nu=1.0),
Matern( d, range=r3, nu=2.0))
matplot( d, y, type="l", lty=1, lwd=2)
xline( 10)
yline( .1)